 Dictionary for logic and
experimental design
Dictionary for logic and
experimental design  Dictionary for logic and
experimental design
Dictionary for logic and
experimental design This dictionary is being assembled to provide a quick reference and concise definitions of terms encountered in papers, seminars, and discussions of experimental design. In some cases, there may be links to additional pages where a more detailed explanation can be provided, including links to the four main pages of this site. Note that other pages on this site indicate a link leads to this dictionary with an asterisk (*). Suggestions for definitions that should be added to this page (as well as the ways they should be linked to other pages) are appreciated and should be sent to John A Wagner (jawagne@med.cornell.edu). You might also enjoy the Dictionary of Cell & Molecular Biolgy. or the On-line Medical Dictionary.
Jump to: A B C D E F G H I J K L M N O P Q R S T U V W X Y Z
Return to: proteins | cDNAs | antibodies | DNA & genes | logic & exptl design | GPIN home Page
A---------------------------------A --------------------------------- TO TOP
Abzyme: An antibody that has catalytic activity; a catalytic antibody. In 1969, William Jencks proposed that antibodies specific for the transition state of a chemical reaction should have catalytic power. Such catalytic antibodies have been produced using transition state analogs or related compounds as immunogens.
Active zone: The area in presynaptic terminals where the specialized secretory machinery for the focused release of neurotransmitters packaged in vesicles is concentrated (i.e. synapsin and related proteins). Under the electron micrograph these areas are recognized as sites where vesicles seem to be undergoing exocytosis (termed omega figures by Rene Couteaux). It is commonly observed that receptors for neurotransmitters are localized on the postsynaptic neuron in close proximity to active zones. This allows for extremely concentrated neuronal signal conduction, in contrast to endocrine cells which release hormones in a diffuse manner such that many cells are stimulated.
Affinity chromatography takes advantage of specific properties of proteins. The column is packed with a material, such as a substrate, ligand, or antibody, to which only specific proteins can bind. The bound proteins can then be displaced off the column by changing the ionic strength of the solvent.
Agonist - a drug (or endogenous molecule) that can bind to a receptor, thereby activating the receptor to elicit a molecular and/or cellular response.
Ames Test: Developed by Dr. Bruce Ames in 1979, this test allows for the classification of chemicals with regards to their carcinogenicity/mutagenicity (Ames, B.N., 1979. Identifying environmental chemicals causing mutations and cancer. Science 204:587-593).
Briefly, Salmonella which contains a specific mutation in the histidine operon (e.g. insertion, deletion, etc...) which makes the strain histidine(-) is grown in a histidine deficient environment. First, the spontaneous reversion rate, which is quite low, is measured. Then, the reversion rate is measured in the presence of a mutagen being tested. The chemical can then be assigned a specific number with respect to its level of mutagenicity and the specific mutation which the chemical causes (e.g. if it causes deletions, it will only revert the Salmonella strain which contained insertions etc.).
Amino Acids: Alpha amino acids are important metabolic intermediates and they are polymerized to form proteins. The Institute for Molecular Biolgy at Jena provides a description of the properties and structures of the amino acids.
Antagonist - a drug which binds to a receptor thereby preventing a response by an agonist or a natural ligand. A pure antagonist has no intrinsic activity of its own. Antagonism can occur by a number of mechanisms:
antibodies: a protein generated by the immune system that specifically recognizes a site on a macromolecule (usually a protein). see Fc and Fab.
antigens: a molecule that generates an immune response and is recognized by an antibody.
Anti-idiotype antibody: An antibody specific to the variable region of the first antibody to a certain antigen (i.e., an antibody that recognizes the antigen binding domain). This antibody mimics the epitope of the original antigen and represents an internal image of the original antigen. Thus, it can be used as vaccine or a probe for receptors.
anti-peptide antibody: an antibody that has been generated against a peptide. Generally the peptide must be conjugated to a carrier protein to generate an immune response. The binding of the peptide should be blocked by an excess of the free peptide, so this is a test of specificity. Peptides can be synthesized even if the protein containing that sequence is not available.
Antisense DNA (Oligonucleotide Therapeutics): blocking translation by hybridizing cDNA (i.e., antisense DNA) to specified mRNA. RNase H is thought to cleave the hybrid, leading to a more rapid mRNA turnover. One of the major difficulties of this approach is that, with increased concentration of oligonucleotide, the specificity toward the targeted mRNA is lowered, thereby inhibiting other random mRNAs. Frequently stable analogues, rather than authentic DNA, are used.
Apoenzyme: An enzyme with no catalytic activity because it lacks a post-translational modification. An apoenzyme must combine with a coenzyme to form a holoenzyme which is catalytically active.
autonomous replication sequence (ARS): A segment of a DNA molecule needed for the initiation of its replication; generally this site must be recognized and bound by the proteins of the replication sequence. All vectors must contain an ARS to replicate.
autoradiography: a technique in which a radioactive material produces an image of itself on a photographic film or emulsion. Autoradiography is used to produce an image of gles containing radiolabled proteins or nucleic acids. In situ hybridization is used to localize RNAs in a cell. Autoradiography can also be used to quantitate HPLC plates, paper chromatography, or colonies of cells.
Atomic force microscopy (a.k.a., AFM): a three-dimensional imaging tool that can measure structures from the atomic level to micron scale. In the most basic AFM, it measures topography by mechanically moving a sharp probe across the sample. It has been combined with an inverted optical microscope capable of confocal imaging. This microscope enables the operator to use fluorescent markers on a wide variety of biological specimens to determine internal structure to 200 nm resolution and determine surface morphology of the sample to 20 nm resolution while imaging under physiological conditions.
B---------------------------------B --------------------------------- TO TOP
Bacterial artificial chromosomes (a.k.a., BACs, fosmids): A high capacity cloning vectors based on E. coli F plasmid. The size of donor DNA can reach great than 300 kbp, so it is very useful in the analysis of large genomes although it usually generate a low yield of donor DNA for the vector maintained at one or two copies per cell. It is introduced into the host by electroporation.
baculovirus expression: a useful eucaryotic expression system that expresses high levels of protein. The baculovirus is a powerful eukaryotic expression system that can be used to express recombinant proteins at levels as high as 500 mg per liter. Many of the posttranslational modification pathways present in mammalian systems are also utilized in baculovirus, allowing the production of recombinant protein which is antigenically, immunogenically, and functionally similar to the native protein. Recombinant protein expression in the baculovirus system encompasses many of the features of prokaryotic and eukaryotic systems. The baculovirus expression system can be used to produce large amounts of target protein in insect host cells. It offers a number of advantages over prokaryotic, yeast, and mammalian expression systems. More than 300 different proteins have been expressed using baculovirus vectors, and in most cases the protein produced is similar in structure, biological activity, and immunological reactivity to the naturally occurring protein..
A baculovirus expression system can use the baculovirus Autographa californica nuclear polyhedrosis virus to produce recombinant proteins in insect cell cultures. Plasmid vectors are used to clone the target gene and transfer it to the baculovirus genome by homologus recombination inside the insect host cells. A restriction digest of the vector results in linearized viral DNA that is missing an essential portion of the baculovirus genome. The target gene is inserted into a transfer vector that carries the missing fragment so that recombination between the transfer vector and the cut DNA restores the essential element and simultaneously transfers the target gene to the baculovirus genome. Following recombination, a few viral plaques are picked and purified, and the recombinant phenotype is verified. The newly isolated recombinant virus can then be amplified and used to infect insect cell cultures to produce large amounts of the desired protein. Some systems bypass the homologous recombination step and perform ligation of the gene of interest to a bacmid in E. coli. Addition of the gene into the bacmid can be detected by lac z disruption. The bacmid can then be isolated and be used to infect the insect cells which will then be the machinery to produce the product of interest.
beta-mercaptoethanol (a.k.a. 2-mercaptoethanol): reagent which can be used to reduce disulfide bonds in proteins. This can help to separate proteins which are dimers, linked by disulfide bonds. 2-mercaptoethanol is commonly used to prepare proteins for gel electrophoresis, see SDS gels. HO-CH2-CH2-SH
Bifunctional Antibodies (aka, Bispecific antibodies): Antibody in which the two antigen-binding sites are specific for separate epitopes (antigenic determinants). They are produced by molecular genetic techniques, fusion of hybridoma cells, or by chemical crosslinking. They can function as the main mediators of cellular cytotoxicity as well as targeting of drugs, toxins, radiolabeled haptens, and effector cells to diseased tissue, primarily tumors.
Bioinformatics: a combination of information technologies to analyze biological information (including genome sequences, protein sequence, genetic information, RNA sequences, RNA expression data, etc.). Bioinformatics is a rapidly growing and evolving field that is attempting to manage all of this information. Searchable databases that contain an immense amount of genetic and protein information.
Biolistic Gun: See gene gun
Blot:
1. (verb) to transfer macromolecules
from a gel (e.g., agarose or acrylamide) to a solid support (usually
nitrocellulose) or
2. (noun) the solid support (usually nitrocellulose) on which
macromolecules have been immobilized.
Thus, this term is used as both a noun and a verb. There are many
types of blots that are named on the basis of the type of
macromolecule transferred and the way it is detected:
Name Macromolecule on support probe used Southern
(capitalized because it is named after its discoverer)DNA DNA northern RNA DNA western protein antibody southwestern protein DNA far western protein protein (usually 32 P labeled)
Botulinum neurotoxin & toxins: serotype A (BoToxA), a potent neurotoxin, is one of seven antigenically distinct proteins produced from strains of Clostridium botulinum. These serotypes (designated A through G) are the causative agents of botulism, a potentially fatal condition of neuromuscular paralysis. The seven types of BoTox, along with tetanus toxin, comprise the clostridial toxin family. These proteins can be useful experimental as selective agents that can interfere with specific cellular functions.
C---------------------------------C --------------------------------- TO TOP
Calcium-phosphate precipitation: A method up precipitating DNA to prepare it for use in transfection.
CAT: see chloramphenicol acetyltransferase.
catalytic antibody: see abzyme
chloramphenicol acetyltransferase (CAT): a bacterial gene that acetylates and inactivates chloramphenicol; frequently used as a reporter gene.
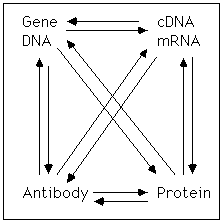
central dogma: The idea that genes, which are sequences of DNA are transcribed to generate a mRNA which can be translated to make proteins.
centimorgan (cM): a unit of distance in a linkage map; specifically, the distance between two linked genes where 1 percent of the products of meiosis are recombinant.
cDNA: a DNA which is complementary to an RNA (a complementary DNA); Generally made by reverse transcription of mRNA.
CDR grafting: replacing of entire complementary determining regions (3 variable loops from the light chain variable domain and 3 loops from heavy chain variable domain) of an antibody by site-directed mutagenesis to reconstruct antibody combining site (ACS) and binding specificity of original antibody while transferring to similar framework.
Chemiluminescence: the emission of light from a chemical reaction. This reaction is of importance in biological research because it is utilized in many mediums to visualize the presence of certain molecules. One of the most common uses is to visualize protein bands on a western blot in place of radioactivity. This is accomplished by using a horse radish peroxidase labeled secondary antibody which causes the emission of photons of light in the presence of a peroxide solution.
Chemokine: small cytokines (6-14kD) that are involved in the migration (chemotaxis) and activation of cells, especially phagocytic cells and lymphocytes. They play a central part in inflammatory response. Chemokines share structural similarities and have been divided into three subfamilies on the basis of the position of conserved cystein residues that form disulfide bonds.
So far, all the known chemokine receptors are heterotrimeric G protein coupled 7 transmembrane proteins. Some of them (CCR5, CXCR4) have been shown to be used as co-receptors for HIV infection.
ChIP (Chromatin immunoprecipitation): a procedure that identifies DNA elements occupied by DNA regulatory proteins in vivo under a given set of conditions. Briefly, proteins are covalently cross-linked to DNA in living cells, the cells are lysed, and DNA is fragmented via sonication. Antibodies to the binding protein can then be used to IP (see immunoprecipitation) the protein-DNA complex. This technique provides a method of purifying the regulatory regions of the DNA bound to the protein at the time of cross-linking. The purified DNA can be amplified and sequence information can be obtained.
Chromatography: Various techniques that are used to separate dissolved components of a mixture as it moves through a porous matrix. Proteins to be fractionated are dissolved in a solvent (mobile phase) and passed through a column (immobile phase). The materials that make up the immobile phase contain sites for specific proteins to bind. As protein molecules bind to the column, their passage is retarded. The greater the affinity of the protein for the material in the matrix, the longer it takes for the protein to move through the column. As the solvent drips out of the end of the column, it is collected into fractions. Proteins with the lowest affinity emerge out of the column first. A description of different types of chromatography. Also see: See ion-exchange chromatography, hydrophobic chromatography, gel filtration chromatography and affinity chromatography. For more information on chrmatography, see this site.
Chromosome walking: a method to assemble long sequences of chromosomal DNA. It involves hybridizing a primer of known sequence to a clone from an unordered genomic library and synthesizing a short complementary strand. The complementary strand is then sequenced and its end used as the next primer for further walking; in this way the adjacent, previously unknown, region is identified and sequenced. The chromosome is thus systematically sequenced from one end to the other. Because primers must be synthesized chemically, a disadvantage of this technique is the large number of different primers needed to walk a long distance. Chromosome walking is also used to locate specific genes by sequencing the chromosomal segments between markers that flank the gene of interest.
Chromosome jumping: a technique, related to chromosome walking, of isolating clones from a genomic library that are not contiguous by skipping a region between known points on the chromosome. Chromosome is done usually to bypass regions that are difficult or impossible to walk through or regions known not to be of interest.
circular dichroism (CD) : an ultraviolet light based spectroscopic technique for determining the relative content of secondary structural features in a protein (alpha-helix, beta-sheet or random coil). This technique takes advantage of the fact that different secondary conformations can be distinguished by their ability to change the ellipticity of light (see web tutorial on CD spectrocopy). Ellipticity is changed because of a preference that one secondary structure has relative to another for absorbing right or left handed light. Ellipticity is the experimental observable and is measured with respect to the wavelength of light impinging upon the detector. Data analysis is performed by least squares fitting to a linear combination of ellipticity versus wavelength curves that correspond to well-characterized secondary structures. CD spectroscopy is in some ways analogous to absorption spectroscopy. Differential absorbance of two different components of light is the measured quantity.
cis-acting: a DNA sequence that acts to control the expression of the gene adjacent to it. In contrast, a trans-acting element acts to regulate the expression of a gene at a distance. Thus, a cis-acting sequence is generally a DNA element in a promoter while a trans-acting sequence codes or regulates the expression of a transcription factor.
Co-transfection: the simultaneous introduction of different plasmids into a given cell. For example, a plasmid expressing an interesting gene promoter) may be introduced along with another plasmid containing a reporter gene, which can be easily assayed or used for drug selection. Often, the reporter gene is used to measure transfection efficiency in experiments to study promoters, since a cell that takes up one type of plasmid will generally take up both. See transfection.
Cold-sensitive: a mutant that exhibita a normal phenotype at the permissive (higher) temperature, but exhibit a mutant phenotype at the restrictive (lower) temperature. Like temperature sensitive mutants, this type of mutation make it possible for the timing of switch from normal to mutant phenotype to be controlled by the experimenter.
Computational neuroscience: the use of knowledge of nervous system physiology (often based on the original Hodgkin-Huxley equations), with the aim of understanding the computations performed by neural systems, and to develop formal models of these computations. Modeling approaches in computational neuroscience generally fall into one of two categories: mathematical models or artificial neural networks. Mathematical models use differential equations to describe the state of an individual neuron, but can be computationally challenging to scale up to neuronal populations. Artificial neural networks use assemblies of elements to model neuronal populations, but sacrifice features of real brain networks, such as synaptic transmission and temporal coding.
confocal laser scanning microscope (CLSM) allows us to observe microscopic structures within thick (tens or hundreds of microns) specimens. With normal fluorescence microscopy, one cannot resolve deep structures within a specimen because of light emitted and scattered by the out-of-focus tissue. By placing small apertures in the light path at points confocal to the focal point within the specimen, light that comes from out-of-focus regions is rejected, allowing detection of just the point of interest. By scanning the light source, a laser beam, across the tissue in the X and Y directions, a high-resolution image of a thin slice of the tissue can be constructed digitally within seconds. From a series of such optical `slices' taken at different depths that are stored in a computer, the three-dimensional representation of the tissue can be generated and manipulated with image-processing software. CLSM has recently become the instrument of choice for biological research, chemical analysis, and materials testing. A few examples.
cM: see centimorgan
Contig Map (short for contiguous map): A genetic map which describes a complete chromosomal segment, usually very large, by the combinations of smaller, contiguous segments of DNA clones which overlap one another:
complementary DNA: a DNA which is complementary to an RNA (a complementary DNA). Generally made by reverse transcription of mRNA. Usually called cDNA.
Complementation test: A genetic test to determine whether two mutations of a given phenotype are in the same genetic unit (i.e. gene) or in separate units (i.e. two different genes). To determine this, genomes containing two recessive mutants are combined in a cell and evaluated for their ability to rescue the mutant phenotype. If the two mutations are in separate genes, each mutant will carry a wild-type copy of the other mutation, thus providing compensation of the defective product. These two mutations are COMPLEMENTARY to each other and exist in different complementation groups. If the two mutations are within the same gene however, there will be no functional wild-type copy of the mutant gene, and therefore these two mutations are assumed to reside in the same complementation group or gene. (Note that mutants within the same gene that can complement one another (intragenetic complementation) do exist, but they are unusual). Complementation should not be confused with recombination.
Cre: an enzyme that catalyzes recombination between lox sites
Cre-loxP recombination: a site specific recombination system that can be designed to function is selective cell types. It is illustrated here. Here is a helpful web site that discusses it.
Chimeric DNA: DNA molecules containing sequences from unrelated genes. Combining coding sequences from two different genes can produce a third protein with novel properties. For example, combining the activator sequences from VP16 with the DNA binding regions of the tet-repressor produces the tTA, a protein that can bind to DNA and activate transcription.
Cytokine: A cytokine is an extracellular signaling protein or peptide that mainly mediates local interactions in cell-cell communication. Many cytokines, especially interleukins and interferons, are secreted by immune cells and are recognized by cytokine receptors on other immune cells. Cytokines cause a variety of actions, such as activation, proliferation, and maturation of the cells. Because cytokines can have many effects on cells, they can be useful in cell culture systems to allow a particular cell type to grow in vitro under physiologically similar conditions. They also allow a researcher to grow a certain type of cell independently that is usually dependent upon another cell for survival. The researcher can simply provide the necessary cytokine(s). Particular cytokines can be added to culture media or removed from the media to determine which signals (cytokines) are necessary for a certain population of cells to survive, to proliferate, or to express differentiated functions. ELISA assays are useful for determining the amount of cytokine that cells are secreting, but measures of cell function (e.g., proliferation) can also provide a sensitive assay for cytokines.
D---------------------------------D --------------------------------- TO TOP
Denaturing Gradient Gel Electrophoresis* (DGGE): Electrophoresis of DNA in an increasing gradient of denaturing agent. When a segment of DNA enters a region where part of the duplex is unstable, the conformation changes and the mobility decreases dramatically. Thus, this method can frequently detect small differences in DNA sequence and is very sensitive to mismatch between two strands. It is frequently used to search for mutations in regions that have been linked to a phenotype. For related methods see 'Screening Methods For Detection Of Unknown Point Mutations.'
DGGE: see Denaturing Gradient Gel Electrophoresis
Diagonal Electrophoresis: Sequential separation of a protein (or another macromolecule) in two directions with some type of reaction to change the mobility of some of the components between the first and second dimension. Components that are insensitive to the reaction run with the same mobility in both directions, so they lie on the diagonal. In contrast, components that are sensitive to the reaction will change mobility during the reaction and will run off the diagonal. Possible reactions include exposing proteins to phosphatase or glyosidases or a chemical that reverses a cross-linking reaction. One use of this approach, which is described below, helps identify the position of disulfide bonds:
DNA arrays (or DNA microarrays): a solid phase systems that allow the simultaneous hybridization, detection, and quantification of multiple (thousands to hundreds of thousands) of nucleic acid sequences. See the 'Genome Chip' site or the 'comparative gene expression site' for details. There are two variants of the DNA microarray technology: 1) cDNA probe (500~5,000 bases long) is immobilized to a solid surface such as glass - using robot spotting and exposed to a set of targets either separately or in a mixture. 2) An array of oligonucleotide (20~25-mer oligos) or peptide nucleic acid probes is synthesized either in situ (on-chip) or by conventional synthesis followed by on-chip immobilization. The array is exposed to labeled sample DNA, hybridized, and the identity/abundance of complementary sequences are determined. Variations of this technology can help study the pattern of gene expression under different conditions or determine the presence of specific genetic variants in populations (for mapping studies) or tissues (to profile tumors).
dominant negative: a phrase used to describe a protein that interferes in a dominant way with the action of a normal protein. e.g., expression of the N17 mutant of ras interferes with the normal signaling by ras.
E---------------------------------E --------------------------------- TO TOP
Electrophoresis: Separation of molecules (frequently macromolecules) by placing them in an electric field. To prevent convection, the molecules must move in some type of stabilized support. Frequently, this is a gel of acrylamide or agarose (depending on the size of the molecule; thus, gel electrophoresis), although electrophoresis can also be done in a gradient (e.g., sucrose gradient) or on a solid support (e.g., paper). Proteins are usually separated on acrylamide gels, frequently after they have been denatured by SDS and beta mercaptoethanol (see SDS-PAGE) but they can also be electrophoresised under native conditions (see native gels). DNA and RNA are more frequently separated on agarose gels (see northern analysis or Southern analysis).
Electroporation : introduction of DNA into cells by passing a high voltage current.
ELISA: See enzyme linked immunoadsorbent assay, below
ELISPOT assay: a variation of the sandwich ELISA* assay which measures the presence of cytokine- or antigen- secreting cells. It is carried out by plating cells on a surface of a plate that has been coated with antibody to a cytokine or other antigen. Cytokine released by the cells can be trapped on the antibody coated plate. Once the cells are washed away, a second antibody can be added to measure the amount of cytokine that is bound to the primary antibody. The ELISPOT assay can be used, for example, on T-cells, to detect cytokine-secreting cells, or on B-cells, to detect specific B-cell antibody secretion.
Embryonic stem cell (ES): a cell line that can participate in all parts of embryonic development, giving rise to all types of differentiated cells, including germ cells. These lines are used as models for differentiation. When mutations are expressed in these cells (a selected mutation, a transgene, or a homologous recombination), the cell can be used to produce and adult animal. These cells can also form tumors (embryonal carcinoma) when injected into an animal.
Endonuclease: Enzyme that cleaves phosphodiester bonds within a nucleic acid chain.
Enhancer: A regulatory DNA sequence that:
Enhancer Trap: A DNA element that has been modified to contain a reporter gene (frequently include the lacZ gene from E.coli) which lacks any promoter can be used to detect enhancers. DNA elements that can integrate into different chromosome sites (insertional mutagenesis). Usually the reporter is not expressed because it lacks a promoter, but tissue- and stage-specific expression of galactosidase from the lacZ gene can be triggered by local signals from enhancer elements adjacent to the insert site. Genes normally influenced by these enhancer elements can be identified by staining for beta-galactosidase activity with X-gal. Thus, this approach serves as a direct way of identifying genes with an interesting expression pattern.
enzyme-linked immunoabsorbent assay (ELISA): an assay (illustrated here) in which the abundance of an antigen (e.g., a protein) is measured by the binding of an antibody, which in turn is determined by an associated enzyme activity. The enzyme which is measured is sometimes cross-linked to the primary antibody, but is more often linked to a 'secondary antibody' which recognizes the primary antibody. The enzyme (frequently HRP (horse radish peroxidase) or a phosphatase) is chosen for technical reasons (low background, stability, easy detection). To be meaningful the assay must be done under conditions where there is an increase in signal with increasing antigen. A good site from the U of Arizona (it even has animation!). For more details.
epitope: a region on a macromolecule which is recognized by an antibody. Frequently it is in a short region of primary sequence in a protein and it is generally about 5 to 12 amino acids long (the size of the antigen binding site on an antibody). Carbohydrates, nucleic acids and other macromolecules may be antigens and have epitopes.
epitope tag: an epitope (see above) is often added to the coding information of an expressed protein to provide a convenient antigenic site that can be recognized by a well characterized antibody. See FLAG-tag
equilibrium density sedimentation: a method of separating cellular components on the basis of their buoyant density, which, unlike sedimentation velocity, is independent of their size and shape. The sample is usually sedimented through a density gradient that contains a very high concentration of sucrose or cesium chloride. Each cellular component begins to move down the gradient, but eventually reaches a position where the density of the solution is equal to its own density. A series of distinct bands is thereby produced in the centrifuge tube, with the bands closest to the bottom of the tube containing the components of highest buoyant density. This method is so sensitive that is capable of separating macromolecules that have incorporated heavy isotopes, such as 13C or 15N, from the same macromolecules that have not. If layered at the bottom of a density gradient, light components can, of course sediment up (i.e., they float). Note contrast to 'velosity sedimentation.'
ESTs (Expressed Sequence Tag): The sequence of a short region of a cDNA (i.e., expressed) that serves to 'tag' the cDNA. Databases of these tags are useful in identifying cDNA, mapping genes, identifying new genes and obtaining clones containing the cDNA, reducing or eliminating the effort required to isolate cDNAs. Because ESTs are partial single-pass DNA sequences (usually 200 ~ 500 bp) reversely transcribed from the 3' or 5' end of mRNA, ESTs represent a sample of coding intron-free sequences of genes. The NCBI-Expressed Sequence Tags Database can be found at http://www.ncbi.nlm.nih.gov/dbEST
Excised patch recordings: see patch-clamp
Exconuclease: a nuclease that degrades nucleic acids by cleaving successive nucleotide residues (or short oligonucleotides) from ends of the strands. 5' and 3' exonucleases are known.
Exonuclease III protection: A method of identifying the site of a protein DNA interaction. This method require labeling a DNA strand at one of the 5' ends. Next, the DNA binding protein is added, followed by the exonuclease. This removes bases from the 3' end, but has to stop when it comes to the binding protein. Removing the proteins from the DNA leaves free bases and sections of DNA labeled at the 5' end, so you can find. The other end is the 3' end of DNA binding site.
expression libraries: a library in which members of the library are expressed and detected by virtue of the translated protein, e.g., see functional cloning of ligand gated channels.
F---------------------------------F --------------------------------- TO TOP
Fab: The region of an antibody that is responsible for binding an antigen (epitope) binding. When it is cleaved from an antibody, which is multivalent, it becomes a monovalent fragment, including parts of two peptide chains. This region of the antibody is characterized by extreme variability that occurs during the production of a gene to be expressed by a B-cell. This variability accounts for the ability different antibodies (there may be many millions of different antibodies) to bind so many different antigens.
FACS: Fluorescence-activated cell sorting (see below).
Fc: The constant region of an antibody. This region, which is in the stem of the antibody is very similar if not identical between different antibodies. Thus, it is a target for secondary antibodies and allows one to detect the binding of the many different antibodies present in an antisera with a single secondary antisera.
FISH (fluorescent in situ hybridization): hybridization of a probe to intact chromosomes to detect the position of a gene. For more information and a diagram see this site.
Fingerprinting: a technique for direct RNA analysis developed by F. Sanger and colleagues. It creates a two-dimensional array of ribonuclease-resistant oligonucleotides. Generally, RNA is digested with RNases including RNase T1 or RNase A. Then, the oligonucleotides are typically separated on the basis of charge in the first dimension and on the basis of size in the second.
FLAG-tag: one of the first epitope tag systems. The FLAG epitope is recognized by commercially available M1 and M2 antibodies in a Calcium dependent binding. The system can be used both for affinity purification and other immunological procedures. more on FLAG-tag
Fluorescence: The property of a molecule that allows it to produce a specific emission wavelength when excited with different (excitation) wavelength of light of higher energy (longer wavelength).
Fluorescence-activated cell sorting: a technique (also called FACS) for analyzing and sorting cells according to their content of fluorescent molecules. A stream of cells is divided into droplets, each containing no more then one cell. A charge is imposed on the surface of the droplets to enable them to be deflected.The droplets are passed one at a time through a focused laser beam, which excites the fluorescence of probe molecules inside the cells or on their surface. Droplets are deflected into different test tubes according to the intensity and spectral distribution of their fluorescence.
Fluorescence anisotropy: This technique is used to study the binding/interaction characteristics between proteins and nucleotide sequences (example: protein/protein interaction, protein/mRNA interaction, protein/DNA interaction). The target molecule is tagged with a fluorescent molecule whose electrons can be excited via a laser. As the excited electron of the fluorescent molecule returns to its ground state energy level, energy will be emitted at a specific wavelength. When polarized light is used, only those molecules in solution aligned with the light source will absorb and emit energy. If the flourescently tagged molecule rotates between the time of absorbance and emission, the emitted energy will be depolarized. The depolarization property is called anisotropy. In solution the tagged molecule will have a specific rate of rotation due to thermal energy, and thus will have a specific anisotropy. If a molecule that binds to the tagged molecule is added to solution, the bound form of the tagged molecule will have slower rate of rotation and therefore will produce a lower amount of depolarization. The change in depolarization is related to the binding characteristics between the two molecules. The advantage of this technique over traditional radioligand binding assays is that one can observe the binding characteristics in real-time, with a relatively small amount of protein. Also, this approach can be more easily scaled up to screen large numbers of potential binding partners of a target molecule.
Fluorescence Resonance Energy Transfer: see FRET
Fluorescent Antibody: An immunoglobulin molecule that has been conjugated with a fluorescent molecule, such as fluorescein, rhodamine, Texas red, or Lucifer yellow, so that it produces fluorescence.
Flow Cytometry: A method used to analyze (or, sometimes fractionate) cells based on a particular fraction's light-absorption or fluorescent properties. A cell sample is passed in a narrow stream through a laser beam and an absorbance or fluorescence profile of the sample is measured. In some cases, successive droplets of the stream may be sorted according to their fluorescence by an automated device. Fluorescence can be due to antibodies that recognize specific cell surface molecules, dyes that bind to DNA, substrates that are modified by cellular enzymes, the abundance of calcium, dyes that respond to membrane potential, etc.
Foot print: a technique that determines which areas of a DNA are bound to a protein, leaving a protein footprint. It is accomplished by a combination of DNA binding and exposure of DNA to conditions that lead to cleavage in the absence of a protein to protect the phosphodiester bond.
FRAP (Fluorescence Recovery After Photobleaching): A technique used to determine the rate at which molecules diffuse on a cells surface. Fluorescent molecules can be bleached by an intense beam of light directed to a particular region of the cell through a microscope. The cell 'restores' molecules after they have been destroyed by photobleaching by an active process or by diffusion from another region of the cell. The rate of recovery can be determined by measuring the fluorescent intensity in the bleached area, and, thus, the number of fluorescent molecules in the area.
FRET (Fluorescence Resonance Energy Transfer): a method used to determine when two macromolecules are in proximity to one another. Most commonly, biologists use this technique to study protein-protein interactions, either in vitro or in vivo. To use this approach, two macromolecule must be labled with a fluorescent tag. Often fusion proteins of the protein of interest with Yellow or Green Fluorescent Protein (GFP or YFP) are generated for this purpose. In this example, GFP is excited by a certain wavelength of light and then emits a green wavelength of light. Subsequently, YFP is excited by a green wavelength of light and emits a yellow wavelength. This energy transfer occurs only if the molecules are in close proximity. When GFP and YFP are not in close proximity, excitation of GFP results in the emision of a green wavelength. When GFP and YFP are in close proximity, excitation of GFP will still result in GFP emitting green light, but this light will be quenched by YFP. As a result, YFP will become excited and emit a yellow wavelength of light. This change in emission spectra allows the experimenter to determine whether two proteins are closely associated or if their association changes under different conditions.
free radical: an atom or group of atoms possessing an unpaired electron. Free radicals include the ROIs and RNIs. They are very reactive and potentially very damaging in biological systems, but they are generated as part of normal metabolic processes and are important and destructive parts of the inflammatory response.
Fusion protein /Chimeric Protein: proteins produced using recombinant DNA techniques; short amino acid sequences of a protein of interest are fused to an easily detected reporter protein so that this sequence can be followed in a cell. One application: most nuclear proteins contain one or more specific short sequences of amino acids that serve as signals for their import into the nucleus after their synthesis in the cytosol. By artificially attaching different segments of such a nuclear protein to a cytoplasmic protein using gene-fusion techniques, the "signal peptides" responsible for nuclear import can be identified. See Chimeric DNA*
G---------------------------------G --------------------------------- TO TOP
G418 sulfate: This antibiotic (aka geneticin) interfers with the function of the large ribosomal subunit and is therefore toxic to bacteria, yeast, protozoa, helminths, and mammalian cells. Resistance can be conferred by the neo gene (neomycin phosphotransferase II ) which is bacterial in origin. This gene can be expressed in eukaryotic cells, allowing positive selection to occur while cells without the neo gene are killed by G418. Cells continue to divide once or twice even in lethal doses of G418, so it may take a few days for effects to become apparent in cell cultures.
Gel filtration chromatography: a chromatography technique that allows separation of proteins on the basis of size. The sample is applied to a column consisting of porous beads made of an insoluble but highly hydrated polymers such as dextran or agarose or polyacrylamide (also known as sizing columns). Sephadex, Sepharose, and Bio-gel are commonly used commercial preparation of these beads, which are typically 0.1mm in diameter. Small molecules can enter these beads, but large ones cannot. The result is that large molecules flow rapidly through this column and emerge first because a smaller volume is accessible to them. In this way proteins can be efficiently separated according to their size (or, more accurately, their stokes radius).
Gel shift analysis: a technique that identifies DNA protein interactions because free oligonucleotides migrate munch faster on a gel than oligonucleotides bound to a protein. Demonstration of specific binding requires evidence that the interaction is high affinity and specific for a unique sequence.
gene disruption: any procedure used to add genetic material to a gene, thereby influencing the expression or function of the gene. The disruption may eliminate function, alter the pattern of expression, or alter the nature of the products of the gene. See knock-in, knock-out, diagram of gene disruption by homologous recombination
Genechip: A miniature device containing short sequences of defined DNA sequence attached to a solid support. These sequences can be designed to test for the expression of particular RNAs or the presence of particular mutant genes or important alleles.
Gene Gun/Biolistic Gun: a mechanical device for introducing foreign genes into cells, particularly those of plants. The device frequently fires a tiny metal particle with a DNA fragment attached to it into a cell.
Gene Trap: An molecular approach to identify genes that are expressed in specific tissues. The principle of gene trap that random integration of a reporter/selector cassette into a genome can both mutate (i.e., by creating an insertion) and identify the trapped locus (i.e., the insertion serves as a genetic marker). In some cases, the presence of splice donor or acceptor elements can lead to the generation of a fusion protein, even in case of integration into an intron, and a reporter protein is expressed in a cell type specific manner. Interesting patterns can be selected for further study. See the Gene Trap Home web site for more information (http://cmhd.mshri.on.ca/sub/genetrap/paradigm.htm)
genomic clone: a cloned DNA sequence that is isolated from the genome.
genomic library: a library consisting of isolated fragments of a genome. It would include introns, exons, non-coding regions, repetitive DNA, as well as structural elements.
H---------------------------------H --------------------------------- TO TOP
HAT selection: a 'positive' selection that allows only cells expressing HGPRTase to survive. The key enzymes using this selection and the way HAT selection is used in making hybridomas can be found at these links.
HGPRTase: the enzyme hypoxanthine-guanine phosphoribosyltransferase. By using these nucleosides as precursors for purines, it bypasses the de novo pathway of purine biosynthesis which can be blocked by aminopterine. It is possible to select against HGPRTase expression with the agent 6-TG.
homologous recombination: a recombination event that occurs between DNAs with the same sequence. Homologous recombination is the mechanism that allows insertion of a targeting vectors into selected (homologous) sequences in procedures that are used to disrupt (knock out) specific (targeted) genes.
Homology screening or cloning: a process to identify new proteins when the sequence encoding one or more related proteins are known. Using sequence information, one can design probes to the (hopefully) conserved regions of the putative family of proteins. Then, one can screen a c-DNA library and clone protein that have related sequences. Analysis of the new cDNA can indicate if it is indeed related to the the original protein. Additional studies of the novel protein(s) can determine if it can perform a new or unknown functions.
HPLC (High Performance Liquid Chromatography): a high resolution separation process using a liquid mobile phase and a column containing microparticulate solid particles coated with a specific functional group. The functional group, which can be neutral, charged, or hydrophobic, cause separation of components of a mixture by the specific physical interaction. The primary modes of HPLC for biological macromolecules are reversed phase, ion exchange, size exclusion, and hydrophobic interaction chromatography. These rapid and high resolution methods have provided a means of purification, separation, and analysis of peptides and proteins.
Hybridoma: a cell line used in the production of monoclonal antibodies*. It is obtained by fusing antibody-secreting B lymphocytes with cells of a lymphocyte tumor.
Hydrophobic Chromatography: chromatography using columns are packed with beads linked to hydrophobic side chains that will bind proteins with exposed hydrophobic regions. Hydrophilic or charged proteins will elute from the column.
Hydrophobicity plots: A plot which is a 'running average' of the hydrophobic (or hydrophylic) nature of a region of a protein. It can be an indication of a the transmembrane helices in proteins, or it can indicate which regions of the protein are buried in the protein away from the solvent, or it can indicate a region of contact between two proteins.
I---------------------------------I --------------------------------- TO TOP
ICAT (Isotope-Coded Affinity Tag): In this approach the proteins present in two samples are labeled separately on the side chains of their reduced cysteine residues using one of two isotopically different ICAT reagents. The samples are mixed, enzymatically digested, subjected to avidin affinity column to isolate peptides labeled with isotope-coded tagging reagents (which contains biotin), and analyzed by MS. Relative quantification of protein expression between the two samples is accomplished by comparison of peak intensities of the isotopically different peptides, and identification is accomplished by selecting these peptides for MS/MS and subsequent sequence database searching with the generated CID spectra. The ICAT reagent consists of three general elements: a reactive group capable of labeling a defined amino acid side chain (e.g. iodoacetamide to modify cysteine residues), an isotopically coded linker, and a biotin tag for the affinity isolation of labeled peptides:

A course at Davidson College has an animation of this approach as well as a proteomics & methods page.
Immunohistochemsitry: the visualization of antigens in their normal cellular and tissue environment. The primary antibody, which detects the antigen of interest is generally detected by a secondary antibody, which is linked to an enzyme (e.g., horse-raddish peroxidase or alkaline phosphatase), a chromogen (e.g., FITC, Fast Red, etc.), or, in the case of electron microscopy, an electron dense material (a gold particle). Use of multiple antibodies and distinguishable detection systems can allow the study of multiple antigens in the same tissue. For more details on immunohistochemistry or immunofluorescence.
in silica: literally, 'in silicon'; this term refores to database serches or computer simulations that are carried out in a computer rather than in a traditional laboratory. See our Bioinformatics page.
in situ: literally 'in place', this designation refers to an assay that is done without disruption of cells or tissue. The term is frequently used to mean in situ hybridization, which detects the presence of a mRNA by hybridization, but also see FISH
in situ hybridization: hybridization of a cDNA or an RNA to fixed tissue which has been treated to make the endogenous RNAs available for hybridization. The tissue must be fixed, dilapidated, treated with proteases, hybridized, and washed to remove non-specifically bound RNA. The hybridized RNA or DNA is usually identified by virtue of label incorporated during synthesis.
in vivo: literally, 'in life'; this term refers to biological studies done in living cells or living organisms.
in vitro: literally 'in glass'; this term refers to studies done in purified systems or, occasionally, in broken cell preparations. Working in vitro is one of the classic pillars of biochemistry and enzymology.
in vivtro: a made up term that is not in common use. It is a combination of in vivo and in vitro. It is designed to indicate studies of pure proteins which are introduced into cells by injection or by transfection. In studies of this type, the entire cell and all its components, are used by the experimentalist as an experimental milieu. Classic examples of this approach include the studies of transcription factors that can interact with the transcription machinery of an intact cell (which is hard to recreate in vitro) or studies of proteins that regulate the cytoskeleton (e.g., rac or rho) and, thus, cell morphology.
insertional mutagenesis: mutagenesis that occurs because of the insertion of an exogenous DNA sequence into a gene. Insertion can occur either by non-specific or specific mechanisms. Recombination at a region of homology with a targeting vector can lead to an intentional disruption of a specific gene, creating a mutation.
internal ribosome entry site: see IRES
immortalizing oncogene: a gene that enables a primary cell to grow (i.e., divide) indefinitely in culture. Cell lines that grow indefinitely in tissue culture must express an immortalizing oncogene. Immortalizing oncogenes can be created by mutation of specific cellular genes (proto-oncogenes) or introduced into a cell by transfection or infection with a virus carrying the gene (e.g., the sarcoma virus). Acquisition of an immortalizing oncogene is one of the steps required for development of a tumor.
immunoprecipitations: The use of an antibody to selectively bind to and precipitate an antigen. Originally this technique relied on the ability of divalent antibodies to cross link and precipitate an antigen. Now, the antigen-antibody complex can be removed from solution by binding it to a Protein A bead. Protein A, which was originally derived from Staph A bacteria, recognizes the Fc region of the antibody. This technique can be effectively combined with SDS gels to visualize the protein or to determine if the precipitated protein has another epitope, like a phoshpo-tyrosine residue or a sugar using western analysis. Likewise, an antibody to one protein will sometimes precipitate another protein bound to the antigen, providing initial evidence for protein-protein interaction.
Ion-exchange chromatography depends on the ionic bonding of a protein to a charged material in the immobile ion-exchange resin. The protein mixture moves through the column in a buffer whose pH promotes the binding of some of the proteins to the resin. The overall charge of a protein depends on the charges of the amino acids that make up the protein. In turn, the charges of the amino acids depend on the pH of the medium. At the isoelectric point, the pH of the medium allows the protein to be neutral, but charged regions of a protein can allow it to bind to the resin even at its isoelectric point. Those molecules bound to the column can then be sequentially displaced by increasing the ionic strength and/or the pH of the buffer.
IRES: internal ribosome entry site are internal sequence on transcribed mRNA where the 40S ribosomal subunit is recruited to mediate cap-independent eukaryotic translation without requiring complexes in the 5` untranslated region (ie. cap). An IRES allows two proteins to be translated from a single mRNA, so they are a useful tool in designing expression vectors. These sequences were originally obtained from retroviruses, which require them in their normal life cycle. Several proteins have been identified for predominantly using IRES sites for translation and can be found at the following website: http://www.rangueil.inserm.fr/IRESdatabase/ .
Isocandamers: restriction enzymes that recognize different DNA sequences but produce same cohesive ends to facilitate cloning.
Isochizomers: restriction enzymes that recognize the same DNA sequence but cut DNA in different ways because they are purified from different organisms.
isoelectric focusing (IEF): a protein purification technique that takes advantage of the unique isoelectric point (i.e., pI) of each protein. A pH gradient is established on a gel or in a gradient using ampholytes (a mixture of small molecules with different pIs). Because of their zwitterionic character, ampholytes migrate when the electric field is applied. Depending on its charge, each ampholine will move towards either the positive or the negative electrode. As they move, a pH gradient is established and each ampholyte will stop moving when it reaches its pI. (Every ampholyte is a buffer.) Likewise, as the pH gradient is established, the protein moves towards the electrode of opposite charge. As the protein(s) migrate through the gradient, they will lose or gain protons until it achieves no net charge, causing it to slow down and stop. It is now at its pI. The pI is the specific pH at which a protein (or ampholyte) has no net charge. Therefore, proteins with a positive charge will loose protons and negatively charged proteins will gain protons to reach this point. If the protein migrates past this point, it will regain its charge and migrate back.
Note that IEF can be used in conjunction with SDS-PAGE to obtain a 2-D gel, where the protein is partially purified first by isoelectric point and then size.
Isoelectric point: the pH where a macromolecule (usually a protein protein) is not charged and, thus, would not move in an electric field and is less likely to bind to an ion-exchange column.
J---------------------------------J --------------------------------- TO TOP
K---------------------------------K --------------------------------- TO TOP
Klenow fragment: the carboxy terminal fragment of DNA Polymerase I, which possesses polymerase activity but lacks exonuclease activity.
knock-in: a variation of gene targeting that uses homologous recombination but allows expression of added genetic sequences in place of the endogenous gene. This approach allows the test of more subtle mutations than is allowed by a simple knock out.
knock-out: 1. a procedure to disrupt the sequence of a gene and interfere with its expression (see diagram). The procedure relies on both homologous recombination and genetic selection (to pick those recombinants where the desired event has occurred). 2. an animal or cell line where a gene has been disrupted. 3. a verb indicating the intent to use the above procedure.
L---------------------------------L --------------------------------- TO TOP
Laemmli gels: an SDS gel with a discontinuous buffer system. It is named for the person who successfully combined these two approaches in 1969.
library: a collection of different macromolecules in a self replicating system. Types include a cDNA library, a genomic library, a phage display library, an expression library.
Ligation of DNA fragments: is most easily accomplished if the two DNA sequences have complementary 'sticky' ends. In this case, the DNA of interest can be incubated with DNA ligase from bacteriophage T4 at temperature and salt where the short duplex is stable.
linkage disequilibrium: the study of closely linked markers in unrelated families (i.e., families whose relationship is so distant that there is no evidence of how they are related) to test for the linkage of very closely linked markers. This approach can frequently rule out some the possibility that some genes are responsible for a particular trait.
linker scanning: a mutagenesis procedure in which short sequences of a promoter are substituted by restrictions sites, usually using a PCR based mutagenesis approach.
lipofectin: a detergent that is used to increase the efficiency of transfection.
LOD score (log of odds): a statistical indication of the probability that a particular trait is linked to a genomic marker. a value of 3 is considered to be strongly significant.
lox: a DNA sequence that is recognized by Cre recombinase, allowing recombination to occur
Luciferase: a gene originally isolated from fire fly that emits a photon in the presence of luciferin and ATP. Production of photons can easily be monitored by a scintillation counter especially designed for this purpose. Luciferase is used as a reporter gene.
lysogeny: one of two possible pathways followed after infection by certain types of bacteriophage, with bacteriophage lambda being the most common example. When following the lysogenic pathway, the phage integrates into the host chromosome where it can remain and replicate along with the host chromosome. It can also excise itself from the host chromosome, replicate, produce structural proteins for the virus, make viruses and produce a burst of progeny. When grown under conditions where a virus follows a lytic pathway, it can form plaques (a clone of the virus).
M---------------------------------M --------------------------------- TO TOP
MALDI (Matrix Assisted Laser Desorption Ionization): An ionization technique in which sample molecules mixed in a UV or IR energy-absorbing matrix are desorbed and ionized by a short laser pulse. MALDI mass spectrometry allows the detection of molecules with masses of several 100 KDa, in combination with time-of-flight mass analyzers. A simplified schematic of MALDI-TOF mass spectrometry is here. A web introduction to MALDI-TOF is here. And see who won the 2002 Nobel Prize in chemistry!
Mass Spectrometer: An instrument used to determine the unique mass spectrum of a compound/chemical. All mass spectrometers are composed of :
The requirements to obtain a mass spectrum are to produce ions in the gas phase, to accelerate these ions to a specific velocity employing electric fields, to seperate the ions into different masses, to send them into a suitable mass analyser, fand to detect each charged entity of a particular mass sequentially in time. see http://hyperphysics.phy-astr.gsu.edu/hbase/magnetic/maspec.html
Maxam Gilbert sequencing: a method for determining the nucleotide sequence of a DNA fragment. The fragment is labeled at one end and then broken preferentially at one of the four nucleotide, under conditions where only an average of about one break is made per fragment. The fragments are separated by gel electrophoresis to find the length of each fragment, which gives the position of each occurrence of the particular nucleotide (ATCG). This procedure is followed using cleavage procedures that give information about each of the 4 nucleotides, yielding the entire sequence of the original fragment. A Maxam Gilbert Sequencing animated diagram is available here.
MCA: monoclonal antibody, see below
methylation interference: a method to identify bases in DNA that make important contacts with DNA binding proteins. First, an oligonucleotide that is thought to bind the protein of interest is radiolabeled at one end. Next, the guanosines and adenosines of the DNA are methylated with dimethylsulfate (DMS) in such a way that the DNA averages one modified site per molecule (i.e., most are not modified). The methylated probe is then incubated with the protein of interest and protein-DNA complexes are formed (but not on sequences where an essential base has been modified). Then, the protein-DNA complex and unbound DNA are separated on a nondenaturing polyacrylamide gel and viewed by autoradiography. In some cases, methylation of a base interferes with the ability of a protein to bind at that base. Therefore, the protein-DNA portion is depleted of any DNA that is methylated at the binding site. Both portions of probe (bound and free) are cleaved specifically at the modified bases with piperidine. They then are analyzed on a denaturing polyacrylamide gel (as in footprint analysis). Lanes derived from free DNA and bound DNA are compared. As in footprinting, bands representing regions of the DNA where the protein bound will be missing in the bound lane yet appear in the free DNA lane. Uracil interference is a similar method that identifies thymines that are involved in protein binding. In this case, the probe is generated via PCR with TTP and dUTP, to incorporate deoxyuracil instead of thymine, and the bound and free bands are cleaved with uracil N glycosylase and piperidine.
Molecular beacon: hairpin-shaped molecules with fluorophore which can be quenched molecule attached to another region of the molecule (see figure). They are designed so that the fluorescence is restored when the beacon hybridizes to a target nucleic acid, but not when the molecule is in its native state. Molecular beacons can be combined with fluorescent spectroscopy for real time monitoring of PCR reactions, the replication cycle of many viruses, or related reactions.
monoclonal antibodies: an antibody preparation which is the product of a cloned gene or a clonal population of cells (generally B-cell hybridomas). Unlike antiserum, which has many different antibodies, all the antibodies in a monoclonal are identical. A MCA (monoclonal antibody) is a well characterized preparation; and, because it comes from a transformed cell type that grows indefinitely in culture, it can be produced in large amounts and be shared my investigators with confidence. In contrast every preparation of antisera, even from the same animal, is different. There is a danger of using a MCAs because an epitope may be shared by different macromolecules, so a MCA may recognize two very different antigens, an important consideration in experimental design.
N---------------------------------N --------------------------------- TO TOP
Native gel: a method of fractionating proteins on the basis of size and shape. In contrast to SDS gels*, the protein is not denatured, so both the shape of the protein and its charge (at the pH where fractionation occurs), contribute to its mobility. Because it is not denatured, as is done before SDS gels*, the protein is more likely to retain biological activity. It may be able to bind small ligands (depending on affinity/off rate) or other proteins, allowing for a semi-quantitative assay.
Neo or Neomycin resistant: see G418.
Nick translation: a method of radioactively labeling double stranded DNA. Nicks on one strand of the DNA are introduced by the endonuclease, deoxyribonuclease I (DNase I). The enzyme DNA polymerase I is then used to add nucleotide residues to the 3' end of the nick. DNA polymerase I also has 5' - 3' exonuclease activity, and this eliminates nucleotides from the 5' end of the nick. Polymerase I thus progresses along one strand on the duplex, incorporating radioactively labeled nucleotides as it does so.
northern analysis: A method to quantitate RNA using a DNA probe. Isolated mRNAs are separated on an agarose or a polyacrylamide gels which are then transferred to nitrocellulose. The transferred mRNAs are then detected by hybridization to a labeled cDNA probe. RNAs are separated by weight so different forms can be measured at the same time. The quality and stability of RNA is always a problem because RNA is so sensitive to degradation, so it is always essential to show that the RNA is of good quality by probing the blot with a control probe that recognized mRNA (looking at tRNA or rRNA is not adequate).
Nuclear Magnetic Resonance (NMR): a procedure which can provide information about the structure of molecules and macromolecules in solution. The basis of this procedure is that certain atomic nuclei (e.g., hydrogen) can generate a magnetic moment, which can take either of 2 orientations when an external magnetic field is applied. Nuclei in different environments (i.e., with different chemical neighbors) absorb energy at slightly different resonance frequencies. This effect is called chemical shift and this shift is expressed in parts per million (ppm). Two-dimensional NMR can provide enough information to solve the structures of peptides and proteins up to the 30 kD range. The Basics of NMR will provide more details and the mathematics of the proceedure is discussed in this course from Dr. Edison at U of Florida.
Nuclear Run On/Run Off Transcription Assay: This assay is used to measure RNA polymerase and transcriptional activity. A nuclear run-on assay is intended to identify a population of labeled RNAs that were being transcribed immediately before isolation. Nuclei from the cells of interest are isolated under conditions where the transcription complexes that have been initiated by RNA polymerase are stable. Run-on reactions are done in the presence of labeled nucleotides (usually 32P-XTPs (in the alpha position)) to label the nascent transcript (i.e., to extend the nascent chain with labeled nucleotides). To identify specific transcript, a dot blot or a slot blot can be done where known DNA samples are spotted on a filter and are allowed to hybridize with the labeled run-on RNA transcripts. A stronger signal suggests that more active polymerase complexes are on the gene of interest.
O---------------------------------O --------------------------------- TO TOP
One-hybrid screening: A powerful method to rapidly identify heterologous transcription factors that can interact with a specific regulatory DNA sequence of interest (the bait sequence). In the one-hybrid system, detection is based on the interaction of a transcription factor (prey) with a bait DNA sequence upstream of a reporter gene. cDNA expression libraries are used to produce hybrids between the prey and a strong trans-activating domain. The advantage of cloning transcription factors or other DNA-binding proteins via one-hybrid screenings, compared to biochemical techniques, is that the procedure does not require specific optimization of in vitro conditions. In this approach, yeast is being used as a "eukaryotic test tube" to demonstrate interactions. This may circumvent possible difficulties, e.g., incorrect protein folding or lack of post-transcriptional modifications, which may occur in screenings for transcription factors via a prokaryotic expression system such as that used in the E. coli-based southwestern procedure. One-hybrid screens are closely related to the two-hybrid screens where the interaction between two proteins (bait and prey) is detected via in vivo reconstitution of a transcriptional activator that turns on expression of a reporter gene.
Orthologue: an evolutionarily related gene from a different species that carries out corresponding functions.
Ouchterlony Gel Diffusion: A largely outmoded method that measures the secondary interaction between antigen and antibody, which is illustrated here. (A secondary interaction is one in which the binding of antibody to antigen causes clumping or precipitation rather than a direct, primary interaction). Antibody and antigen are put in different wells that have been punched into a gel 9usually agarose). They diffuse towards one another and precipitate where they meet at 'equivalence', forming a visible band. When serum samples in two wells are tested agains the same well of antigen, the relatedness of the sameples can be determined by the shape of the precipitin line formed. This approach has been largely superceeded by western analysis*.
ORF: open reading frame
open reading frame (ORF): an uninterrupted series of triplets that does not contain stop codons; long ORFs presumably encode a protein.
Osmotic Mini Pump: Osmotic pumps are plastic containers planted in subjects (frequently subcutaneously). The pump releases its contents at a controlled rate for the life of the pump. The pump works with an osmotic gradient that fills one of its chambers, thereby squeezing its innermost chamber, which is filled with infusate. Alzet is a commercial supplier.
P---------------------------------P --------------------------------- TO TOP
PAGE: polyacrylamide gel electrophoresis; see SDS gels and Electrophoresis
Paralogue: An evolutionarily related family of genes within one species that were derived from a common ancestor. Paralogues have presumably diverged to carry out similar but divergent functions.
Patch-clamp Technique: a relatively new (1976) microelectrode technique that can record current flow across a cell membrane. This method is improved over traditional intracellular recording because it can isolate (both physically and electrically) a small number of channels. In the patch-clamp technique, a small glass microelectrode with a tip diameter of approximately 1 micron (a larger tip than intracellular electrodes) is pressed against the plasma membrane of a cell to form a high resistance seal. Generally, the voltage across the 'patch' of membrane is 'voltage clamped*' (hence the name 'patch-clamp') to a known voltage, allowing the currents flowing across the membrane to be measured. Using this approach, it is possible to see the current flowing through a single channel. i.e., it is possible to see a single channel opening and then closing in real time. This is one of the few situations where the activity of a single molecule can be quantitatively studied. Since it is possible to express mutant channels, it is possible to do detailed structure function studies of There are a number of variations that can be done with related techniques:
The whole cell variant of the patch clamp technique (on-cell recording and whole cell recording, above) allow recording from , respectively, a patch of membrane on the an intact cell or an intact cell. Excised patch recordings allow recording of conductance across a small fragment of the membrane, but depending on how it is made, the excised patch can be in either orientation.
P1 element: a vector that can carry inserts that are tens of kbases long and are very useful in genomic cloning.
PCR: see polymerase chain reaction
PCR-Assisted Binding Site Selection: A method used to determine the sequence of DNA target regions bound by proteins with unknown DNA-binding specificity. This is a general method of detecting DNA-protein interactions. The general protocol is as follows and here is a diagram of the procedure:
Phage Display Library: a selection technique in which a peptide or protein is expressed as a fusion with a coat protein of a bacteriophage, resulting in display of the fused protein on the exterior surface of the phage particle, while the DNA encoding the fusion resides within the virion. The affinity of the phage for a macromolecule of interest is a rapid way to identify specific interactions.
Plasmid: a small circular DNA molecule that is distinct from the bacterial genome and replicates autonomously. Plasmids, pieces of plasmids, and plasmids with specific modifications can be used to introduce foreign genetic information into cells, conferring antibiotic resistance or the expression of specific proteins, reporters or RNAs.
Plaque purification: The isolation of a genetically pure virus by prorogation of a group of viruses at a dilution large enough that a plaque arising from a single virus can be isolated.
For example, plaque purification is used to isolate a single recombinant phage plaque from a library through serial dilutions of a large plug (containing progeny from several independent infections) from a portion of the library which gives a signal when hybridized with appropriate probe(s). Dilutions of phage from such a plug are used to transfect bacteria, which are mixed with top agarose and plated on bottom agar. Lifts are performed on sparsely populated plates and hybridized with probe. From the autoradiogram, an individual phage colony (i.e., a plaque) can be isolated by using the small end of a Pasteur pipette. For discussion see discussion in 'cDNA cloning'.
This procedure is analogous to purifying bacteria by isolating a single colony (i.e., the bacteria that are derived from a single 'founder').
Polyadenylation: the final step during the formation of a mature mRNA, the addition of a series of adenosine residues to the 3' end of the mRNA. Many eucaryotic primary transcripts are cleaved by a specific endonuclease that recognizes the AAUAAA poly(A) signal, then a poly(A) polymerase adds about 250 A residues to the 3' end of the transcript. Polyadenylation is very important for mRNA stability, mRNA nuclear-cytoplasmic transport, and translation into functional proteins.
polyclonal antibodies: a preparation of antibodies derived from multiple clones of B-cells created during an immune response. An antiserum contains polyclonal antibodies, along with many other serum proteins.
polyacrylamide gel: see SDS gels
polymerase chain reaction or PCR: a series of reactions that result in the amplification of a sequence between two primers (or, in advanced modifications of PCR, between a primer and a DNA end (RACE)). It can be adapted for many purposes including screening libraries, measuring RNA abundance or localization, looking for novel transcripts, or introducing sequences (mutations) into DNA constructs. A mini-animation of PCR is on the Cold Spring Harbor learning site.
Polytene Chromosome: In Dipteran flies (e.g. Drosophila and Chironomous), these large chromosomes are easily visualized during the interphase stage of the cell cycle. They consist of a large number of partially replicated chromosomes neatly stuck together in a lateral array. These chromosomes, housed in the larval salivary glands, are easily and reproducibly stained and viewed under a light microscope, so their banding pattern was among the first physical markers of DNA. They were also the first chromosomes used for FISH*. These tightly compacted chromosomes reveal a characteristic decondensation when a gene is being expressed resulting in a distinctive "puff".
primer: a short nucleic acid sequence (an oligonucleotide) which can hybridize to a long sequence and be extended by a polymerase. Sometimes a primer is chosen to be specific (e.g., primer extension), but sometimes random primers are used to prime all the possible nucleic acids in a mix (e.e., to produce a labeled probe).
Prion (proteinaceous infectious particle): A proposed pathogen composed only of protein with no detectable nucleic acid and which is responsible a series of genetic, sporadic or infectious neurodegenerative disorders in humans (Kuru, Creutzfeldt-Jakob disease, Gerstmann-Straussler-Scheinker disease, and fatal familiar insomnia) and animals (scrapie, bovine spongiform encephalopathy). The current hypothesis is that a mutant protein has the ability to affect the folding of its homologue in other cells or organisms to create another folded mutant which can then disrupt other proteins like itself. This method of infection is still controversial because many scientists feel that there must be a genetic component to transmit the disease.
primer extension: a technique in which an oligonucleotide primer (which has been hybridized to an longer nucleic acid) is extended with a polymerase to determine the sequence of the the nucleic acid 5' of the oligo. This technique is used to map transcription initiation sites (i.e., the most 5' end of a mRNA).
probe: a labeled oligonucleotide or nucleic acid used to detect others by hybridization. Probes are used in many techniques including northern analysis, screening libraries, doing RNAase protection assays, etc.
Promoter: a nucleotide sequence in DNA recognized by RNA polymerase and transcription factors to allow initiation of transcription.
Protein Crystals: A crystal is composed of an object, or motif, repeated translationally on a three-dimensional lattice. Protein crystals can be grown from a supersaturated solution of a pure protein. The supersaturation state is usually achieved by lowering the protein solubility through addition of a precipitating agent, which is most commonly a salt (e.g., ammonium sulfate, sodium chloride, sodium citrate), and organic solvent (e.g., ethanol, methylpentanediol, acetone), or a polyethylene glycol. Other factors that affect protein solubility include protein concentration, pH, temperature, and the presence of specific ligands and metal ions. The identification of conditions appropriate for protein crystal growth is largely a multidimensional search by trial and error, and often it represents the major obstacle in a crystallographic analysis.
Proteome: A catalog of all proteins coded by the genome of an organism. Originally, the term was taken to imply a static list of all proteins coded by the genome. More recently researchers have employed a more dynamic definition of proteome as the identity of all proteins, their chemical modifications the way those proteins interact, and their 3 dimensional structure.
proteomics: a systematic study of the functions, interactions, cellular location, expression and post-translational modifications of proteins on a massively parallel scale. Some of the approaches to this set of cutting edge are introduced on this proteomics site.
purification table: A table that includes information about the fractionation of material during a purification. It records the amount of material present in each fraction containing the biological activity of interest, the level of the biological activity that is being measured, and the specific activity of the fraction (activity divided by amount of material), the fold purification, and the yield at each step. Analysis of this table indicates the success and the quality of a purification scheme. Although most commonly used to monitor protein purification, it can also be used to monitor the purification of any substance.
Q---------------------------------Q --------------------------------- TO TOP
Quantitative real-time RT-PCR: A method of mRNA quantification that involves the design of specific PCR primers and a fluorescently labeled probe that anneals to the PCR product. The largest advantage to this technique is the ability to monitor the PCR reaction in real-time, ensuring that PCR reaction efficiencies are equal between PCR products, and that quantification is based on the linear portion of the reaction, where a back calculation to starting mRNA is possible. A common way to do this is described below:
R---------------------------------R --------------------------------- TO TOP
RACE: acronym for Rapid Amplification of cDNA Ends. RACE is a technique that can be used to identify unknown sequences at the 5' or 3' ends of DNA transcripts. It is an application based on the principles of RT-PCR, but uses one sequence specific primer and one additional primer that anneals to either the poly A tail of mRNA's (this is 3'RACE) or a tail added to cDNA ends (5'RACE). Commonly, the sequence information gathered from RACE reactions are used to generate new sequence specific primers that can then amplify the entire cDNA sequence of interest.
radioimmunoassay: see RIA, below.
Restriction Fragment Length Polymorphism: see RFLP.
RFLP (for Restriction Fragment Length Polymorphism): A variation of Southern analysis which identifies variations between individuals in a population based on the differences in DNA fragment sizes produced from genomic DNA by particular restriction endonucleases. Specific polymorphic sequences in the genome serve as lineage markers in genetic analysis.
real-time RT-PCR : see Quantitative real-time RT-PCR
reporter: 1. a gene that is used to assay the function of a promoter element. It is usually a gene that is easy to assay. Its expression can be used to test the importance of particular DNA sequences in a promoter or to test the importance of the expression of a transcription factor or a signal transduction pathway on gene expression. 2. Sometimes, the term 'reporter' is generalized to indicate any easy assay that is used to indicate the function of another molecule.
regulatory elements: a DNA sequence present in a promoter that contributes to the regulation of gene expression. a cis-acting DNA sequence, i.e., a DNA sequence that acts to control the gene that is adjacent to it.
Restriction Enzyme (a.k.a. restriction endonuclease): an enzyme that recognizes specific base sequences in double-helical DNA and cleave both strands of the duplex at specific places. Originally discovered in a wide variety of prokaryotes where it cleaves foreign DNA molecules (endogenous DNA is not degrades because the sites recognized by its own restriction enzymes are methylated).
Reverse genetics: jargon for an approach in which a gene is mutated to determine phenotype.
reverse transcriptase (RT): a polymerase that produces a cDNA from an RNA. This enzyme is used to make cDNA libraries and it is essential to the life cycle of retroverses (RNA viruses that replicate by a DNA intermediate).
RIA: Radioimmunoassay uses an antibody antigen reaction to measure antigen concentration. A known amount of labeled antigen is mixed with an unknown and precipitated with antibody under conditions where antibody is limiting. As the amount of unknown antigen increases, less of the labeled antigen can be precipitated. This is somewhat similar to an ELISA assay, but the measurement is based on a competition rather than a direct recognition of the antigen. The antigen is traditionally labeled with an isotope, but other, non-radioactive methods of detecting the antigen can be used in the same way. For more information.
Ribozyme: an RNA molecule that can break or form covalent bonds in their own sequence another molecule. i.e., it is capable of acting as an enzyme. The reactions observed include cleaving themselves or other RNA molecules, ligation, and trans-splicing. Ribozymes greatly accelerate the rate of the reaction, and can show extraordinary specificity with respect to the substrates it acts on and the products it produces. There are three type o f ribozymes:
RNAase protection: a method of quantitating RNA based on its ability to form a nuclease resistant hybrid with a labeled probe. The more RNA, the more probe can be protected. If only part of the probe hybridizes to the RNA of interest (i.e., the probe has 5' or 3' regions that are not homologous to the RNA of interest), then only part of the probe is protected. The protected probe and the intact probe will fractionate at different places on a gel. Protection of a fragment of a unique and predictable size is good criteria for specificity and in established assays, quantitation can also be done by precipitating the protected probe. The probe can be either an RNA or a DNA probe, but RNA probes have the advantage of being more stable at high temperatures and they can be made to a higher specific activity using RNA polymerase.
RNAi, see short inhibitory RNAs
RNI: reactive nitrogen intermediate including a nitrogen atom, e.g., NO. (nitric oxide).
ROI: reactive oxygen intermediat including an oxygen atom, e.g., OH. (hydroxyl radical).
RT-PCR: an abbreviation for a method of detecting RNA by initially transcribing the RNA with reverse transcriptase (RT), and then amplifying the cDNA with the Polymerase Chain Reaction (PCR). This can be done analytically, preparatively, or in order to determine where an RNA is synthesized. This is a sensitive technique, but controls to establish if the reaction is specific or linear (quantitative) are essential.
S---------------------------------S --------------------------------- TO TOP
SAGE (Serial Analysis of Gene Expression): a powerful technique for the analysis of mRNA abundance in different cell types, having the same purpose, but different from DNA chip technology since it doesn't involve hybridization. In SAGE a short (close to 9bp) unique sequence is used as a particular mRNA tag. This tag is the 9 base pairs immediately to the right of a four base restriction site located closest to the 3' end of a message. It can either have to do with known or unknown mRNA sequence. SAGE involves ligating groups of these tags together and sequencing them. The abundance of a mRNA is related to the number of times a specific tag appears compared to the total number of tags sequenced. This technique is an efficient approach, which lets you analyze many mRNAs at the same time. It estimates mRNA abundance for both known and unknown genes.
- Cartoon to assist with understanding the concept:
________________CATGxxxxxxxxx________________AAAAAA� 3' end of cDNA
________________GTACxxxxxxxxx________________TTTTTTT�
4 BRS* 9 bp tag
Sanger sequencing: A method of sequencing DNA through controlled interruption of replication. DNA polymerase is used to replicate a particular sequence, but an analog (2', 3' deoxyXTP) of one of the bases is included in the incubation mixture, along with normal copies of all four bases. Replication stops when the analog base is used, so the replicated sequence is always blocked at a particular base. The process is repeated with deoxy analogues of all four bases. Gel electrophoresis is used to determine the lengths of the fragments. This procedure allows a rapid determination of the entire sequence.
Sedimentation Coefficient ('S value'): The rate at which each component sediments depends primarily on its size, shape and density. Measurements of sedimentation coefficients are routinely used to help determine the size and subunit composition of the organized assemblies of the macromolecules found in cells. The sedimentation coefficients of biological macromolecules are expressed in Svedberg units (abbrev. as S; 1S= 1.0 x 10exp-13 seconds. ) Named after Svedberg, the Swedish inventor of the ultracentrifuge.
short inhibitory RNAs (siRNA or RNAi): 19-21 nucleotide sequences of RNA that, when introduced into cells, guide endogenous RNA degradation proteins to complementary mRNA sequences specifically initiating degradation and preventing synthesis. By reducing RNA levels, siRNA can interfere with protein synthesis and can be used analogously to genetic knockouts to study the role of a specific protein. For a diagram of the mechanism, click here.
siRNA, see short inhibitory RNAs
SSCP: Single-Strand Conformational Polymorphism, below.
Single-Strand Conformational Polymorphism (SSCP): electrophoretic separation of ss nucleic acid based on subtle differences in sequence (often a single base pair) which results a different secondary structure and a measurable difference in mobility. This method provides a mechanism to rapidly screen DNA for mutations. For additional discussion of this and related methods see 'Screening Methods For Detection Of Unknown Point Mutations.'
Site-directed mutagenesis: mutagenesis in which one or a few bases are changed. Site directed mutagenesis in DNA is useful for analyzing promoter and enhancer sequences, and for studying gene regulation. In protein (i.e., mutation of coding sequences in DNA) it can be used to address role of a particular region/motif/domain/residues of the protein, or study the effect of naturally occurring mutations on protein function. Common methods of creating mutations are the Kunkel mutagenesis and the PCR-based mutagenesis. See defining cis acting sequences
SDS gels: a method of fractionating proteins according to size (+/-10%). Proteins are denatured with SDS, a strong detergent and a reducing agent, like DTT. The SDS gives the proteins a negative charge proportional to their weight and so strong that it generally eliminates the effect of intrinsic charge. The proteins move thorough a gel of acrylamide (chosen because it does not interact strongly with proteins) under the influence of an electric field (see electrophoresis). Smaller proteins have a smaller frictional coefficient and move faster. Proteins can be fixed to the gel and visualized with CBB (Coomassie Brilliant Blue) which can detect as little as 0.1 micrograms of protein. Alternatively, proteins can be transferred to nitrocellulose and detected with antibodies or other reagents (see western blots). Current systems use a buffer system including three different pHs and different ions which act to focus the proteins to a narrow band before fractionation begins, increasing resolution. The combination of discontinuous buffers and SDS was introduced by Laemmli in 1969, so these are commonly called Laemmli gels.
SDS-PAGE: abreviation for SDS Polyacrylamide gel electrophoresis. see SDS gels, electrophoresis.
site-specific recombinases: an enzyme that promotes recombination between specific DNA sequences.
in situ: literally 'in place', this designation refers to an assay that is done without disruption of cells or tissue. The term is frequently used to mean in situ hybridization, which detects the presence of a mRNA by hybridization, but also see FISH
in situ hybridization: hybridization of a cDNA or an RNA to fixed tissue which has been treated to make the endogenous RNAs available for hybridization. The tissue must be fixed, dilapidated, treated with proteases, hybridized, and washed to remove non-specifically bound RNA. The hybridized RNA or DNA is usually identified by virtue of label incorporated during synthesis.
sizing column: see gel filtration chromatography
small nuclear RNA (snRNA): a group of polynucleotides, usually from 70 to 500 nucleotides in length. As complexes with proteins (snrps) they have several functions, among them the processing of pre-mRNA either directly as part of a spliceosome or indirectly by hybridizing with the pre-mRNA and altering its architecture. Some snrps are encoded by a separate gene while some are found within introns.
Single Nucleotide Polymorphism (SNPs): single base pair positions in genomic DNA at which normal individuals in a given population show different sequence alternatives (alleles) with the least frequent allele having an abundance of 1 % or greater. SNPs occur once every 100 to 300 bases, hence, they are the most common genetic variations.
Southern analysis: Detection of DNA by its hybridization to DNA after it has been transferred to nitrocellulose. The technique of northern blotting was named because of its similarity to Southern Blotting, which was named after its originator, and , thus, should be capitalized. A mini-animation of Southern analysis is on the Cold Spring Harbor learning site.
Southwestern blot: Detection of protein by its hybridization to a labeled DNA probe. Used to look at DNA-protein interactions (i.e. transcriprion factors).
SSCP: see single-strand conformational polymorphism,above
Stable transfection: introduction of a DNA into a cells and genetic selection of a cell line that expresses the gene.
Subtractive hybridization: A technique used to enrich for particular mRNAs present in one cell but not another. Closely related cells are used, only one of which produces a particular protein (or proteins) of interest. cDNA is synthesized from the cell that makes the protein, and this cDNA is hybridized with mRNA from the 2nd cell. cDNA sequences that remain unpaired are enriched for ones that encode for the protein. These sequences can be purified from the soluble stranded nucleic acid by hydroxyapitite chromatography, a technique that separates single-stranded and double-stranded nucleic acids.
surface plasmon resonance biosensor: an optical biosensor that is a useful tool for the qualitative and quantitative characterization of the specific binding of a mobile reactant to a binding partner immobilized on a sensor surface.
Synaptosome preparation: A homogenization and fraction method that produces a preparation that is enriched in 'pinched off' nerve terminals. Brain (different areas depending on the study), are homogenized then fractionated by differential centrifugation and sucrose density gradients. The resulting membrane fractions are not anatomically distinguishable neurons, but rather, a preparation that is enriched in a membrane compartment which functions, in some ways, as a nerve terminal. One can conduct studies of neurotransmitter release upon stimulation by different drugs, or just by changing extracellular ion concentrations and thus changing membrane potential which will alter membrane permeability (e.g. through voltage gated channels). One can then examine which neurotransmitters are released, the relative quantity of each, any changes in neurotransmitter release upon drug addition and ions involved in causing the release. Neurotransmitters can also be added to the extracellular solution and see how they affect their own release or release of other neurotransmitter. Cautions should be exercised because these preparations are always heterogeneous and because they synaptosomes are always heavily contaminated with other membranes that can complicate results.
T---------------------------------T --------------------------------- TO TOP
"T-bag" technique: a method for simultaneous parallel solid-phase synthesis of either combinatorial libraries or compound arrays. It involves the use of a porous, flexible mesh pouch (or tea bag) that is filled with resin beads and exposed to the desired reagents at various stages of the solid-phase synthesis process. This method also provides a simple method for removal of excess reagents and impurities. A library produced by this method can be screened directly after cleavage from the resin, or it can be used to create another new library. For example, a peptide library composed of more than 24 million hexapeptides with acetylated N terminals and amidated C terminals can be made by this process. The first two positions are individually defined, however the last four positions consist of equimolar mixtures of 18 and 20 natural L-amino acids. These mixtures can then be screened for a desired activity, and then the last four positions can be defined by an iterative process (each stage reduces the number of peptide sequences by 20-fold). This tea bag technique has allowed for the use of combinatorial chemistry in drug discovery. The technique was patented by Dr. Richard Houghten.
Telomerase: a specialized eukaryotic DNA polymerase with both RNA and protein components. It regulates the structure of chromosome ends by adding variable numbers of telomeric simple sequence repeats. This de novo addition is thought to be required to compensate for the loss of terminal repeats that occurs with each round of genome replication. In turn, the number of telomeric simple sequence repeats regulates chromosome stability, gene expression, cell proliferation, and possibly other events as well.
Temperature-sensitive / ts: a mutant that exhibit a normal phenotype at the permissive (lower) temperature, but exhibit a mutant phenotype at the restrictive (higher) temperature. (i.e., it is heat-sensitive). Also see cold sensitive. Like cold sensitive mutants, this type of mutations make it possible for the timing of switch from normal to mutant phenotype to be controlled by the experimenter.
TOF (Time-of-Flight): A TOF analyzer measures the flight times of ions; lighter ions traverse a given distance faster than heavier ions. The mass is proportional to the time squared. Flight times are converted to m/z values by calibration with compounds giving ions of known m/z values. A simplified schematic of MALDI-TOF mass spectrometry is here. An definition of MALDI is provided above. A web introduction to MALDI-TOF is here. And see who won the 2002 Nobel Prize in chemistry!
tTA(tetracycline (tet) -regulated trans-activation system, including 'tet-on' and 'tet-off'): a powerful tool for the temporal and quantitative control of exogenous genes expression in mammalian cells and transgenic mice. tTA is a fusion protein composed of the tetracycline repressor domain of E.coli and the transcriptional activation domain (VP16) of herpes simplex virus. There are two versions of the system. In one version, tTA binds to the tetracycline resistance operator (tetO) in the absence of tetracycline. This DNA element is just upstream of the CMV minimal promoter (or another promoter) and the gene of interest. Then the activation domain of tTA activates the transcription of the gene of interest. in the presence of tetracycline, a conformational change in tet repressor of tTA prevents tTA from binding to resistance operator, thus, inhibiting transcription of the gene of interest (i.e., it is a tet-off regulation). Other variants of the hybrid work in the opposite way. In these cases the binding of tet is required for binding to the DNA element and the activation of transcription (so this is a tet on system).
Three hybrid screening: A method to screen for interaction between nucleic acids and proteins using two two-hybrid proteins and a single hybrid nucleic acid that links the hybrid proteins and activates transcription if the hybrid nucleic interacts specifically with a component of each of the hybrid proteins. One of the proteins binds to a promoter and the nucleic acid of interest, and the other binds to the nucleic acid of interest and an activation domain.
transfection efficiency: a measurement of the frequency that a DNA is introduced into a cell during transfection. Transfection efficiency may vary dramatically, so a measurement of transfection efficiency is often essential to interpret data. Data on reporter function is normalized for transfection efficiency using a cotransfected plasmid as a measure of transfection efficiency. see co-transfection
Transient transfection: introduction of a DNA into a cells without any effort to maintain expression
Transition state analogs: molecules that resemble the geometry of the substrate's transition state in the enzyme-substrate complex, and, so, competitively inhibit the substrate's ability to bind to the enzyme. For example, penicillin is an effective transition state analog that uses a moiety similar to that of the natural substrate,and, furthermore, forms a covalent bond with a Ser residue at the active site of glycopeptide transpeptidase, which is the enzyme that is normally involved in the synthesis of peptidoglycan. (Peptidoglycan which is the component of the cells wall that gives the bacteria mechanical support and prevents it from bursting).
Transposons: Discovered in the 1950's, these "movable genes" insert at various points in the genome and often cause pleiotrophic mutations. The simplest bacterial transposon consists of an insertion sequence (IS) region of up to 3,000 base pairs in length, flanked on both sides by inverted repeat regions. The IS region often encodes a transposase enzyme which is involved in the transposition events (some have lost this element). These bacterial transposons jump via DNA intermediates. Eukaryotic transposons include Ac and Ds elements in corn which cause color mutations, HML/HMR mating loci in yeast, P elements in Drosophila, Ty elements in yeast that often activate silent genes, and Alu repeats in animals. The last two examples are representative of retroposons, elements that use transcribed RNA molecules to jump into new locations in the DNA.
Transposable elements are often used in genetic manipulations. They can be utilized in mapping of cloned genes by introducing a plasmid into E. coli that carries a chromosome with a transposon and Amp resistance in it. If a transposon jumps into the plasmid, these plasmids can be isolated because they disrupt the gene carried on a plasmid. Restriction enzymes can be used to map the positions of insertions, and DNA next to the insertions can be sequenced using transposons as primers. Recently, transposable elements began to be used in order to deliver DNA into cells. P element has been used to place promotorless lacZ gene into different points in the genome, so that they adopt expression patterns of various genes (if placed downstream of the transcription regulatory regions with enhancers/promoters). By detecting b-gal activity, one can trace differential expression of genes in different cells.
TUNEL: an acronym for TdT (terminal deoxynucleotidyl transferase)-mediated dUTP nick end labeling. DNA degradation is considered the key biochemical event in apoptosis resulting in cleavage of nuclear DNA into nucleosome-sized fragments (approximately 180 bp unit size). TUNEL can be used to test for the existance of these fragments in individual cells. The principle is that DNA ends generated during apoptosis are extended by TdT using either biotin-labeled or digoxigenin-labeled BrdUrd (bromodeoxyuridine). Incorporated material will be detected by fluorescent reagents. The TUNEL method preferentially labels apoptotic compared to necrotic cells.
Two D gel (Two dimensional gel): a method of protein fractionation that first fractionates a protein mix by isoelectric focusing and then by SDS electrophoresis is illustrated here and here (in more detail)
Two hybrid screening: A method of screening cDNA libraries on the basis of protein-protein interaction (for a diagram look here). Briefly, this approach requires introducing three vectors into a cell:
2-mercaptoethanol: see beta-mercaptoethanol
trans-acting: a DNA sequence that acts to control the expression of the gene that is not closely linked. In contrast, a cis-acting element acts to regulate the expression of adjacent genetic information. Thus, a cis acting sequence is generally a DNA element in a promoter while a trans-acting sequence codes or regulates the expression of a transcription factor.
Transformation:
transgene: a gene that is introduced into a species, such as a mouse, to study the function of the gene product. The gene is generally controlled by a promoter chosen by the experimentalist to allow expression in particular tissues or developmental stages.
targeting vector: a vector designed to recombine into an endogenous gene by homologous recombination. An insertion can disrupt the gene or replace specific sequences depending on the design of the vector.
U---------------------------------U --------------------------------- TO TOP
UV Cross-linking of Protein to Nucleic acid: Cross-linking proteins to nucleic acids with UV light is a simple method for rapidly and accurately determining the molecular weight of a DNA-binding protein in a crude extract. The purpose of the method is to specifically transfer a radio label from a DNA-binding site to the binding protein. Irradiation of DNA with UV light produces purine and pyrimidine free radicals. If a protein molecule is in close proximity to the free radical, a covalent bond can be formed, i.e. cross-linking the protein to the DNA.
UTR: untranslated region- The areas of an mRNA which flank the 'coding' sequence which encodes a protein. UTRs are present at both the 5' end (5'UTR) and the 3' end (3'UTR). UTRs often contains regulatory information about the mRNA's stability, its translation efficiency, and, sometimes, its subcellular localization. See getting full length cDNAs.
URF (unassigned reading frame): An open reading frame (ORF) whose function has not yet been determined.
V---------------------------------V --------------------------------- TO TOP
vector: a self replicating organism used to maintain a library or a piece of DNA of interest.
Velocity sedimentation: a method of separating components of a cellular homogenate or mixture. The homogenate is layered as a narrow band on top of a dilute salt solution that fills a centrifuge tube. When centrifuged, subcellular components sediment at different speeds according to their size and frictional coefficient (which depends on their shape) and their density (which creates a driving force). To stabilize the sedimenting bands against convective mixing caused by small differences in temperature or solute concentration, the tube contains a continuous shallow gradient of sucrose that increases in concentration toward the bottom of the tube (typically from 5%-10% or 10-30% sucrose). Following centrifugation, the different components can be collected individually, most simply by puncturing the plastic centrifuge tube and collecting drops from the bottom. When done under appropriate conditions and properly calibrated it can be used to measure sedimentation coefficient. Note the contrast with equilibrium sedimentation.
Viroid: a plant pathogen that consists of a naked RNA molecule of approximately 250-350 nucleotides, whose extensive base pairing results in a nearly correct double helix.
Virus: an infectious particle composed of a protein capsule and a nucleic acid core, which is dependent on a host organism for replication. While the genome of a DNA virus can replicate directly, a retrovirus must first generate a double-stranded DNA copy from its RNA genome. This DNA is then integrated into the host chromosome and transcription of the integrated DNA can form new RNA viral genomes.
in vivo: literally, 'in life'; this term refers to biological studies done in living cells or living organisms.
In vitro: literally 'in glass'; this term refers to studies done in purified systems or, occasionally, in broken cell preparations. Working in vitro is one of the classic pillars of biochemistry and enzymology.
in vivtro: a made up term that is not in common use. It is a combination of in vivo and in vitro. It is designed to indicate studies of pure proteins which are introduced into cells by injection or by transfection. In studies of this type, the entire cell and all its components, are used by the experimentalist as an experimental milieu. Classic examples of this approach include the studies of transcription factors that can interact with the transcription machinery of an intact cell (which is hard to recreate in vitro) or studies of proteins that regulate the cytoskeleton (e.g., rac or rho) and, thus, cell morphology.
Voltage-Clamp: a technique used to keep the voltage across the membranes constant while measuring changes in current. To achieve this, a circuit which can measure intracellular voltage and inject a current to prevent it from changing even if there is a substantial flow of current across the membrane is created. This is a very important technique in neuropharmacology and membrane physiology, because it allows the experimentalist to monitor changes in current upon application of different drugs to neurons or other cell types. In combination with the use of ion replacement, inhibitors of known ion channels and other toxins, it is possible to establish the involvement of a particular ion channel in the mechanism of a drug or a natural ligand.
W---------------------------------W --------------------------------- TO TOP
western blotting (western analysis). A technique to identify an antigen after fractionation by gel electrophoresis (see SDS gels). Briefly, after electrophoresis, the gel is laid against a sheet of nitrocellulose. The protein is then transferred to the nitrocellulose by electrophoresis (proteins stick to nitrocellulose). After blocking the nitrocellulose (the blot) with high concentrations of nonspecific proteins (usually milk or serum), the antigens are bound with and antibody. The antibody is then detected with a secondary antibody. This secondary antibody can be linked to a easily measured enzyme like HRP (horse radish peroxidase) or the ECL reagent (a reagent that generates light by a chemical reaction). For more information.
X---------------------------------X --------------------------------- TO TOP
x-ray crystallography: a technique for determining the three-dimentional arrangement of atoms in a molecule based on the diffraction pattern of x-rays passing through a crystal of the molecule. This approach is used extensively to solve the structures of proteins and nucleic acids.
Y---------------------------------Y --------------------------------- TO TOP
YACs: yeast artificial chromosomes: a vector that can carry large sequences of genomic DNA (millions of base pairs) and is useful in genomic cloning.
yeast artificial chromosomes: YACs: a vector that can carry large sequences of genomic DNA (millions of base pairs) and is useful in genomic cloning.
Z---------------------------------Z---------------------------------TO TOP
Zymogen /Proenzyme: An inactive precursor to an activated enzyme. Often the zymogen requires a proteolytic cleavage to become active.
ZZZZZZZZZZZZZZZZZZZZZZZZZZZZZ
-------------- ZATSALLFOLKS ![]()
Return to: proteins | cDNAs | antibodies | DNA & genes | bioinformatics | logic & exptl design (home page)| GPIN home Page